Summation theorems (biochemistry)
In metabolic control analysis, a variety of theorems have been discovered and discussed in the literature.[1][2][3][4] The most well known of these are flux and concentration control coefficient summation relationships. These theorems are the result of the stoichiometric structure and mass conservation properties of biochemical networks.[5][6] Equivalent theorems have not been found, for example, in electrical or economic systems.
The summation of the flux and concentration control coefficients were discovered independently by the Kacser/Burns group[7] and the Heinrich/Rapoport group[8] in the early 1970s and late 1960s.
If we define the control coefficients using enzyme concentration, then the summation theorems are written as:
However these theorems depend on the assumption that reaction rates are proportional to enzyme concentration. An alternative way to write the theorems is to use control coefficients that are defined with respect to the local rates which is therefore independent of how rates respond to changes in enzyme concentration:
Although originally derived for simple linear chains of enzyme catalyzed reactions, it became apparent that the theorems applied to pathways of any structure including pathways with complex regulation involving feedback control.[9][10]
Derivation
There are different ways to derive the summation theorems. One is analytical and rigorous using a combination of linear algebra and calculus.[11] The other is less rigorous, but more operational and intuitive. The latter derivation is shown here.
Consider the two-step pathway:
where and are fixed species so that the system can achieve a steady-state.
Let the pathway be at steady-state and imagine increasing the concentration of enzyme, , catalyzing the first step, , by an amount, . The effect of this is to increase the steady-state levels of S and flux, J. Let us now increase the level of by such that the change in S is restored to the original value it had at steady-state.
The net effect of these two changes is by definition, .
There are two ways to look at this thought experiment, from the perspective of the system and from the perspective of local changes. For the system we can compute the overall change in flux or species concentration by adding the two control coefficient terms, thus:
We can also look at what is happening locally at every reaction step for which there will be two: one for , and another for . Since the thought experiment guarantees that , the local equations are quite simple:
where the terms are the elasticities. However, because the enzyme elasticity is equal to one, these reduce to:
Because the pathway is linear, at steady-state, . We can substitute these expressions into the system equations to give:
Note that at steady state the change in and must be the same, therefore .
Setting , we can rewrite the above equations as:
We then conclude through cancelation of since , that:
Interpretation
The summation theorems can be interpreted in various ways. The first is that the influence enzymes have over steady-state fluxes and concentrations is not necessarily concentrated at one location. In the past, control of a pathway was considered to be located at one point only, called the master reaction or rate limiting step. The summation theorem suggests this does not necessarily have to be the case.
The flux summation theorem also suggests that there is a total amount of flux control in a pathway such that if one step gains control another step most lose control.
Although flux control is shared, this doesn't imply that control is evenly distributed. For a large network, the average flux control will, according to the flux summation theorem, be equal to , that is a small number. In order for a biological cell to have any appreciable control over a pathway via changes in gene expression, some concentration of flux control at a small number of sites will be necessary. For example, in mammalian cancer cell lines, it has been shown[12] that flux control is concentrated at four sites: glucose import, hexokinase, phosphofructokinase, and lactate export.
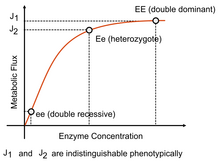
Moreover, Kacser and Burns[13] suggested that since the flux–enzyme relationship is somewhat hyperbolic, and that for most enzymes, the wild-type diploid level of enzyme activity occurs where the curve is reaching a point in the curve where changes have little effect, then since a heterozygote of the wild-type with a null mutant will have half the enzyme activity it will not exhibit a noticeably reduced flux. Therefore, the wild type appears dominant and the mutant recessive because of the system characteristics of a metabolic pathway. Although originally suggested by Sewall Wright,[14][15] the development of metabolic control analysis put the idea on a more sound theoretical footing. The flux summation theorem in particular is consistent with the flux summation theorem for large systems. Not all dominance properties can be explained in this way but it does offers an explanation for dominance at least at the metabolic level.[16]
Concentration summation theorem
In contrast to the flux summation theorem, the concentration summation theorem sums to zero. The implications of this are that some enzymes will cause a given metabolite to increase while others, in order to satisfy the summation to zero, must cause the same metabolite to decrease. This is particularly noticeable in a linear chain of enzyme reactions where, given a metabolite located in the center of the pathway, an increase in expression of any enzyme upstream of the metabolite will cause the metabolite to increase in concentration. In contrast, an increase in expression of any enzyme downstream of the metabolite will cause the given metabolite to decrease in concentration.[17]
See also
References
- ^ Westerhoff, Hans V. (27 May 2023). "Summation Laws in Control of Biochemical Systems". Mathematics. 11 (11): 2473. doi:10.3390/math11112473.
- ^ Sauro, Herbert M.; Small, J. Rankin; Fell, David A. (May 1987). "Metabolic control and its analysis. Extensions to the theory and matrix method". European Journal of Biochemistry. 165 (1): 215–221. doi:10.1111/j.1432-1033.1987.tb11214.x. PMID 3569295.
- ^ Hofmeyr, Jan-Hendrik S.; Kacser, Henrik; Merwe, Kirsten J. (March 1986). "Metabolic control analysis of moiety-conserved cycles". European Journal of Biochemistry. 155 (3): 631–640. doi:10.1111/j.1432-1033.1986.tb09534.x. PMID 3956502.
- ^ Rankin Small, J.; Fell, David A. (January 1989). "The matrix method of metabolic control analysis: its validity for complex pathway structures". Journal of Theoretical Biology. 136 (2): 181–197. Bibcode:1989JThBi.136..181R. doi:10.1016/S0022-5193(89)80225-5. PMID 2779266.
- ^ Schuster, Stefan (October 1996). "Control Analysis in Terms of Generalized Variables Characterizing Metabolic Systems". Journal of Theoretical Biology. 182 (3): 259–268. Bibcode:1996JThBi.182..259S. doi:10.1006/jtbi.1996.0163. PMID 8944157.
- ^ Hofmeyr, Jannie (2001). "Metabolic control analysis in a nutshell". S2CID 17007756.
{{cite journal}}: Cite journal requires|journal=(help) - ^ Kacser, H.; Burns, J. A. (1973). "The control of flux". Symposia of the Society for Experimental Biology. 27: 65–104. PMID 4148886.
- ^ Heinrich, R.; Rapoport, T. A. (1974). "A linear steady-state treatment of enzymatic chains. General properties, control and effector strength". European Journal of Biochemistry. 42 (1): 89–95. doi:10.1111/j.1432-1033.1974.tb03318.x. PMID 4830198.
- ^ Reder, Christine (November 1988). "Metabolic control theory: A structural approach". Journal of Theoretical Biology. 135 (2): 175–201. Bibcode:1988JThBi.135..175R. doi:10.1016/S0022-5193(88)80073-0. PMID 3267767.
- ^ Heinrich R. and Schuster S. (1996) The Regulation of Cellular Systems, Chapman and Hall.
- ^ Heinrich R. and Schuster S. (1996) The Regulation of Cellular Systems, Chapman and Hall.
- ^ Tanner, Lukas Bahati; Goglia, Alexander G.; Wei, Monica H.; Sehgal, Talen; Parsons, Lance R.; Park, Junyoung O.; White, Eileen; Toettcher, Jared E.; Rabinowitz, Joshua D. (July 2018). "Four Key Steps Control Glycolytic Flux in Mammalian Cells". Cell Systems. 7 (1): 49–62.e8. doi:10.1016/j.cels.2018.06.003. PMC 6062487. PMID 29960885.
- ^ Kacser, Henrik; Burns, James A (1 March 1981). "The Molecular Basis of Dominance". Genetics. 97 (3–4): 639–666. doi:10.1093/genetics/97.3-4.639. PMC 1214416. PMID 7297851.
- ^ Wright, Sewall (January 1934). "Physiological and Evolutionary Theories of Dominance". The American Naturalist. 68 (714): 24–53. doi:10.1086/280521. S2CID 84400871.
- ^ Vasseur, François; Fouqueau, Louise; de Vienne, Dominique; Nidelet, Thibault; Violle, Cyrille; Weigel, Detlef (24 April 2019). "Nonlinear phenotypic variation uncovers the emergence of heterosis in Arabidopsis thaliana". PLOS Biology. 17 (4): e3000214. doi:10.1371/journal.pbio.3000214. PMC 6481775. PMID 31017902.
- ^ Billiard, Sylvain; Castric, Vincent; Llaurens, Violaine (December 2021). "The integrative biology of genetic dominance". Biological Reviews. 96 (6): 2925–2942. doi:10.1111/brv.12786. PMC 9292577. PMID 34382317.
- ^ Sauro, Herbert (2013). Systems biology: an introduction to metabolic control analysis (1st, version 1.01 ed.). Seattle, WA: Ambrosius Publishing. ISBN 978-0982477366.

































