Proteinogenic amino acid

Proteinogenic amino acids are amino acids that are incorporated biosynthetically into proteins during translation. The word "proteinogenic" means "protein creating". Throughout known life, there are 22 genetically encoded (proteinogenic) amino acids, 20 in the standard genetic code and an additional 2 (selenocysteine and pyrrolysine) that can be incorporated by special translation mechanisms.[1]
In contrast, non-proteinogenic amino acids are amino acids that are either not incorporated into proteins (like GABA, L-DOPA, or triiodothyronine), misincorporated in place of a genetically encoded amino acid, or not produced directly and in isolation by standard cellular machinery (like hydroxyproline). The latter often results from post-translational modification of proteins. Some non-proteinogenic amino acids are incorporated into nonribosomal peptides which are synthesized by non-ribosomal peptide synthetases.
Both eukaryotes and prokaryotes can incorporate selenocysteine into their proteins via a nucleotide sequence known as a SECIS element, which directs the cell to translate a nearby UGA codon as selenocysteine (UGA is normally a stop codon). In some methanogenic prokaryotes, the UAG codon (normally a stop codon) can also be translated to pyrrolysine.[2]
In eukaryotes, there are only 21 proteinogenic amino acids, the 20 of the standard genetic code, plus selenocysteine. Humans can synthesize 12 of these from each other or from other molecules of intermediary metabolism. The other nine must be consumed (usually as their protein derivatives), and so they are called essential amino acids. The essential amino acids are histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, and valine (i.e. H, I, L, K, M, F, T, W, V).[3]
The proteinogenic amino acids have been found to be related to the set of amino acids that can be recognized by ribozyme autoaminoacylation systems.[4] Thus, non-proteinogenic amino acids would have been excluded by the contingent evolutionary success of nucleotide-based life forms. Other reasons have been offered to explain why certain specific non-proteinogenic amino acids are not generally incorporated into proteins; for example, ornithine and homoserine cyclize against the peptide backbone and fragment the protein with relatively short half-lives, while others are toxic because they can be mistakenly incorporated into proteins, such as the arginine analog canavanine.
The evolutionary selection of certain proteinogenic amino acids from the primordial soup has been suggested to be because of their better incorporation into a polypeptide chain as opposed to non-proteinogenic amino acids.[5]
Structures
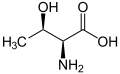
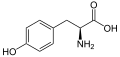
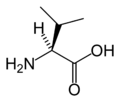
The following illustrates the structures and abbreviations of the 21 amino acids that are directly encoded for protein synthesis by the genetic code of eukaryotes. The structures given below are standard chemical structures, not the typical zwitterion forms that exist in aqueous solutions.

- L-Alanine
(Ala / A) - L-Arginine
(Arg / R) - L-Asparagine
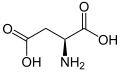
(Asn / N) - L-Aspartic acid
(Asp / D) - L-Cysteine
(Cys / C) - L-Glutamic acid
(Glu / E) - L-Glutamine
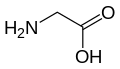
(Gln / Q) - Glycine
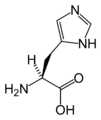
(Gly / G) - L-Histidine
(His / H) - L-Isoleucine
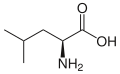
(Ile / I) - L-Leucine
(Leu / L) - L-Lysine
(Lys / K) - L-Methionine
(Met / M) - L-Phenylalanine
(Phe / F) - L-Proline
(Pro / P) - L-Serine
(Ser / S) - L-Threonine
(Thr / T) - L-Tryptophan
(Trp / W) - L-Tyrosine
(Tyr / Y) - L-Valine
(Val / V)
IUPAC/IUBMB now also recommends standard abbreviations for the following two amino acids:
- L-Selenocysteine
(Sec / U) - L-Pyrrolysine
(Pyl / O)
Chemical properties
Following is a table listing the one-letter symbols, the three-letter symbols, and the chemical properties of the side chains of the standard amino acids. The masses listed are based on weighted averages of the elemental isotopes at their natural abundances. Forming a peptide bond results in elimination of a molecule of water. Therefore, the protein's mass is equal to the mass of amino acids the protein is composed of minus 18.01524 Da per peptide bond.
General chemical properties
| Amino acid | Short | Abbrev. | Avg. mass (Da) | pI | pK1 (α-COO-) |
pK2 (α-NH3+) |
|---|---|---|---|---|---|---|
| Alanine | A | Ala | 89.09404 | 6.01 | 2.35 | 9.87 |
| Cysteine | C | Cys | 121.15404 | 5.05 | 1.92 | 10.70 |
| Aspartic acid | D | Asp | 133.10384 | 2.85 | 1.99 | 9.90 |
| Glutamic acid | E | Glu | 147.13074 | 3.15 | 2.10 | 9.47 |
| Phenylalanine | F | Phe | 165.19184 | 5.49 | 2.20 | 9.31 |
| Glycine | G | Gly | 75.06714 | 6.06 | 2.35 | 9.78 |
| Histidine | H | His | 155.15634 | 7.60 | 1.80 | 9.33 |
| Isoleucine | I | Ile | 131.17464 | 6.05 | 2.32 | 9.76 |
| Lysine | K | Lys | 146.18934 | 9.60 | 2.16 | 9.06 |
| Leucine | L | Leu | 131.17464 | 6.01 | 2.33 | 9.74 |
| Methionine | M | Met | 149.20784 | 5.74 | 2.13 | 9.28 |
| Asparagine | N | Asn | 132.11904 | 5.41 | 2.14 | 8.72 |
| Pyrrolysine | O | Pyl | 255.31 | ? | ? | ? |
| Proline | P | Pro | 115.13194 | 6.30 | 1.95 | 10.64 |
| Glutamine | Q | Gln | 146.14594 | 5.65 | 2.17 | 9.13 |
| Arginine | R | Arg | 174.20274 | 10.76 | 1.82 | 8.99 |
| Serine | S | Ser | 105.09344 | 5.68 | 2.19 | 9.21 |
| Threonine | T | Thr | 119.12034 | 5.60 | 2.09 | 9.10 |
| Selenocysteine | U | Sec | 168.053 | 5.47 | 1.91 | 10 |
| Valine | V | Val | 117.14784 | 6.00 | 2.39 | 9.74 |
| Tryptophan | W | Trp | 204.22844 | 5.89 | 2.46 | 9.41 |
| Tyrosine | Y | Tyr | 181.19124 | 5.64 | 2.20 | 9.21 |
Side-chain properties
| Amino acid | Short | Abbrev. | Side chain | Hydro- phobic |
pKa§ | Polar | pH | Small | Tiny | Aromatic or Aliphatic |
van der Waals volume (Å3) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Alanine | A | Ala | -CH3 | - | - | Aliphatic | 67 | ||||
| Cysteine | C | Cys | -CH2SH | 8.55 | acidic | - | 86 | ||||
| Aspartic acid | D | Asp | -CH2COOH | 3.67 | acidic | - | 91 | ||||
| Glutamic acid | E | Glu | -CH2CH2COOH | 4.25 | acidic | - | 109 | ||||
| Phenylalanine | F | Phe | -CH2C6H5 | - | - | Aromatic | 135 | ||||
| Glycine | G | Gly | -H | - | - | - | 48 | ||||
| Histidine | H | His | -CH2-C3H3N2 | 6.54 | weak basic | Aromatic | 118 | ||||
| Isoleucine | I | Ile | -CH(CH3)CH2CH3 | - | - | Aliphatic | 124 | ||||
| Lysine | K | Lys | -(CH2)4NH2 | 10.40 | basic | - | 135 | ||||
| Leucine | L | Leu | -CH2CH(CH3)2 | - | - | Aliphatic | 124 | ||||
| Methionine | M | Met | -CH2CH2SCH3 | - | - | Aliphatic | 124 | ||||
| Asparagine | N | Asn | -CH2CONH2 | - | - | - | 96 | ||||
| Pyrrolysine | O | Pyl | -(CH2)4NHCOC4H5NCH3 | N.D. | weak basic | - | ? | ||||
| Proline | P | Pro | -CH2CH2CH2- | - | - | - | 90 | ||||
| Glutamine | Q | Gln | -CH2CH2CONH2 | - | - | - | 114 | ||||
| Arginine | R | Arg | -(CH2)3NH-C(NH)NH2 | 12.3 | strongly basic | - | 148 | ||||
| Serine | S | Ser | -CH2OH | - | - | - | 73 | ||||
| Threonine | T | Thr | -CH(OH)CH3 | - | - | - | 93 | ||||
| Selenocysteine | U | Sec | -CH2SeH | 5.43 | acidic | - | ? | ||||
| Valine | V | Val | -CH(CH3)2 | - | - | Aliphatic | 105 | ||||
| Tryptophan | W | Trp | -CH2C8H6N | - | - | Aromatic | 163 | ||||
| Tyrosine | Y | Tyr | -CH2-C6H4OH | 9.84 | weak acidic | Aromatic | 141 |
§: Values for Asp, Cys, Glu, His, Lys & Tyr were determined using the amino acid residue placed centrally in an alanine pentapeptide.[6] The value for Arg is from Pace et al. (2009).[7] The value for Sec is from Byun & Kang (2011).[8]
N.D.: The pKa value of Pyrrolysine has not been reported.
Note: The pKa value of an amino-acid residue in a small peptide is typically slightly different when it is inside a protein. Protein pKa calculations are sometimes used to calculate the change in the pKa value of an amino-acid residue in this situation.
Gene expression and biochemistry
| Amino acid | Short | Abbrev. | Codon(s) | Occurrence | Essential‡ in humans | |||
|---|---|---|---|---|---|---|---|---|
| in Archaean proteins (%)& |
in Bacteria proteins (%)& |
in Eukaryote proteins (%)& |
in human proteins (%)& | |||||
| Alanine | A | Ala | GCU, GCC, GCA, GCG | 8.2 | 10.06 | 7.63 | 7.01 | No |
| Cysteine | C | Cys | UGU, UGC | 0.98 | 0.94 | 1.76 | 2.3 | Conditionally |
| Aspartic acid | D | Asp | GAU, GAC | 6.21 | 5.59 | 5.4 | 4.73 | No |
| Glutamic acid | E | Glu | GAA, GAG | 7.69 | 6.15 | 6.42 | 7.09 | Conditionally |
| Phenylalanine | F | Phe | UUU, UUC | 3.86 | 3.89 | 3.87 | 3.65 | Yes |
| Glycine | G | Gly | GGU, GGC, GGA, GGG | 7.58 | 7.76 | 6.33 | 6.58 | Conditionally |
| Histidine | H | His | CAU, CAC | 1.77 | 2.06 | 2.44 | 2.63 | Yes |
| Isoleucine | I | Ile | AUU, AUC, AUA | 7.03 | 5.89 | 5.1 | 4.33 | Yes |
| Lysine | K | Lys | AAA, AAG | 5.27 | 4.68 | 5.64 | 5.72 | Yes |
| Leucine | L | Leu | UUA, UUG, CUU, CUC, CUA, CUG | 9.31 | 10.09 | 9.29 | 9.97 | Yes |
| Methionine | M | Met | AUG | 2.35 | 2.38 | 2.25 | 2.13 | Yes |
| Asparagine | N | Asn | AAU, AAC | 3.68 | 3.58 | 4.28 | 3.58 | No |
| Pyrrolysine | O | Pyl | UAG* | 0 | 0 | 0 | 0 | No |
| Proline | P | Pro | CCU, CCC, CCA, CCG | 4.26 | 4.61 | 5.41 | 6.31 | No |
| Glutamine | Q | Gln | CAA, CAG | 2.38 | 3.58 | 4.21 | 4.77 | No |
| Arginine | R | Arg | CGU, CGC, CGA, CGG, AGA, AGG | 5.51 | 5.88 | 5.71 | 5.64 | Conditionally |
| Serine | S | Ser | UCU, UCC, UCA, UCG, AGU, AGC | 6.17 | 5.85 | 8.34 | 8.33 | No |
| Threonine | T | Thr | ACU, ACC, ACA, ACG | 5.44 | 5.52 | 5.56 | 5.36 | Yes |
| Selenocysteine | U | Sec | UGA** | 0 | 0 | 0 | >0 | No |
| Valine | V | Val | GUU, GUC, GUA, GUG | 7.8 | 7.27 | 6.2 | 5.96 | Yes |
| Tryptophan | W | Trp | UGG | 1.03 | 1.27 | 1.24 | 1.22 | Yes |
| Tyrosine | Y | Tyr | UAU, UAC | 3.35 | 2.94 | 2.87 | 2.66 | Conditionally |
| Stop codon† | - | Term | UAA, UAG, UGA†† | ? | ? | ? | — | — |
* UAG is normally the amber stop codon, but in organisms containing the biological machinery encoded by the pylTSBCD cluster of genes the amino acid pyrrolysine will be incorporated.[9]
** UGA is normally the opal (or umber) stop codon, but encodes selenocysteine if a SECIS element is present.
† The stop codon is not an amino acid, but is included for completeness.
†† UAG and UGA do not always act as stop codons (see above).
‡ An essential amino acid cannot be synthesized in humans and must, therefore, be supplied in the diet. Conditionally essential amino acids are not normally required in the diet, but must be supplied exogenously to specific populations that do not synthesize it in adequate amounts.
& Occurrence of amino acids is based on 135 Archaea, 3775 Bacteria, 614 Eukaryota proteomes and human proteome (21 006 proteins) respectively.[10]
Mass spectrometry
In mass spectrometry of peptides and proteins, knowledge of the masses of the residues is useful. The mass of the peptide or protein is the sum of the residue masses plus the mass of water (Monoisotopic mass = 18.01056 Da; average mass = 18.0153 Da). The residue masses are calculated from the tabulated chemical formulas and atomic weights.[11] In mass spectrometry, ions may also include one or more protons (Monoisotopic mass = 1.00728 Da; average mass* = 1.0074 Da). *Protons cannot have an average mass, this confusingly infers to Deuterons as a valid isotope, but they should be a different species (see Hydron (chemistry))
| Amino acid | Short | Abbrev. | Formula | Mon. mass§ (Da) | Avg. mass (Da) |
|---|---|---|---|---|---|
| Alanine | A | Ala | C3H5NO | 71.03711 | 71.0779 |
| Cysteine | C | Cys | C3H5NOS | 103.00919 | 103.1429 |
| Aspartic acid | D | Asp | C4H5NO3 | 115.02694 | 115.0874 |
| Glutamic acid | E | Glu | C5H7NO3 | 129.04259 | 129.1140 |
| Phenylalanine | F | Phe | C9H9NO | 147.06841 | 147.1739 |
| Glycine | G | Gly | C2H3NO | 57.02146 | 57.0513 |
| Histidine | H | His | C6H7N3O | 137.05891 | 137.1393 |
| Isoleucine | I | Ile | C6H11NO | 113.08406 | 113.1576 |
| Lysine | K | Lys | C6H12N2O | 128.09496 | 128.1723 |
| Leucine | L | Leu | C6H11NO | 113.08406 | 113.1576 |
| Methionine | M | Met | C5H9NOS | 131.04049 | 131.1961 |
| Asparagine | N | Asn | C4H6N2O2 | 114.04293 | 114.1026 |
| Pyrrolysine | O | Pyl | C12H19N3O2 | 237.14773 | 237.2982 |
| Proline | P | Pro | C5H7NO | 97.05276 | 97.1152 |
| Glutamine | Q | Gln | C5H8N2O2 | 128.05858 | 128.1292 |
| Arginine | R | Arg | C6H12N4O | 156.10111 | 156.1857 |
| Serine | S | Ser | C3H5NO2 | 87.03203 | 87.0773 |
| Threonine | T | Thr | C4H7NO2 | 101.04768 | 101.1039 |
| Selenocysteine | U | Sec | C3H5NOSe | 150.95364 | 150.0489 |
| Valine | V | Val | C5H9NO | 99.06841 | 99.1311 |
| Tryptophan | W | Trp | C11H10N2O | 186.07931 | 186.2099 |
| Tyrosine | Y | Tyr | C9H9NO2 | 163.06333 | 163.1733 |
Stoichiometry and metabolic cost in cell
The table below lists the abundance of amino acids in E.coli cells and the metabolic cost (ATP) for synthesis of the amino acids. Negative numbers indicate the metabolic processes are energy favorable and do not cost net ATP of the cell.[12] The abundance of amino acids includes amino acids in free form and in polymerization form (proteins).
| Amino acid | Short | Abbrev. | Abundance (# of molecules (×108) per E. coli cell) |
ATP cost in synthesis | |
|---|---|---|---|---|---|
| Aerobic conditions |
Anaerobic conditions | ||||
| Alanine | A | Ala | 2.9 | -1 | 1 |
| Cysteine | C | Cys | 0.52 | 11 | 15 |
| Aspartic acid | D | Asp | 1.4 | 0 | 2 |
| Glutamic acid | E | Glu | 1.5 | -7 | -1 |
| Phenylalanine | F | Phe | 1.1 | -6 | 2 |
| Glycine | G | Gly | 3.5 | -2 | 2 |
| Histidine | H | His | 0.54 | 1 | 7 |
| Isoleucine | I | Ile | 1.7 | 7 | 11 |
| Lysine | K | Lys | 2.0 | 5 | 9 |
| Leucine | L | Leu | 2.6 | -9 | 1 |
| Methionine | M | Met | 0.88 | 21 | 23 |
| Asparagine | N | Asn | 1.4 | 3 | 5 |
| Pyrrolysine | O | Pyl | - | - | - |
| Proline | P | Pro | 1.3 | -2 | 4 |
| Glutamine | Q | Gln | 1.5 | -6 | 0 |
| Arginine | R | Arg | 1.7 | 5 | 13 |
| Serine | S | Ser | 1.2 | -2 | 2 |
| Threonine | T | Thr | 1.5 | 6 | 8 |
| Selenocysteine | U | Sec | - | - | - |
| Valine | V | Val | 2.4 | -2 | 2 |
| Tryptophan | W | Trp | 0.33 | -7 | 7 |
| Tyrosine | Y | Tyr | 0.79 | -8 | 2 |
Remarks
| Amino acid | Abbrev. | Remarks | |
|---|---|---|---|
| Alanine | A | Ala | Very abundant and very versatile, it is more stiff than glycine, but small enough to pose only small steric limits for the protein conformation. It behaves fairly neutrally, and can be located in both hydrophilic regions on the protein outside and the hydrophobic areas inside. |
| Asparagine or aspartic acid | B | Asx | A placeholder when either amino acid may occupy a position |
| Cysteine | C | Cys | The sulfur atom bonds readily to heavy metal ions. Under oxidizing conditions, two cysteines can join in a disulfide bond to form the amino acid cystine. When cystines are part of a protein, insulin for example, the tertiary structure is stabilized, which makes the protein more resistant to denaturation; therefore, disulfide bonds are common in proteins that have to function in harsh environments including digestive enzymes (e.g., pepsin and chymotrypsin) and structural proteins (e.g., keratin). Disulfides are also found in peptides too small to hold a stable shape on their own (e.g. insulin). |
| Aspartic acid | D | Asp | Asp behaves similarly to glutamic acid, and carries a hydrophilic acidic group with strong negative charge. Usually, it is located on the outer surface of the protein, making it water-soluble. It binds to positively charged molecules and ions, and is often used in enzymes to fix the metal ion. When located inside of the protein, aspartate and glutamate are usually paired with arginine and lysine. |
| Glutamic acid | E | Glu | Glu behaves similarly to aspartic acid, and has a longer, slightly more flexible side chain. |
| Phenylalanine | F | Phe | Essential for humans, phenylalanine, tyrosine, and tryptophan contain a large, rigid aromatic group on the side chain. These are the biggest amino acids. Like isoleucine, leucine, and valine, these are hydrophobic and tend to orient towards the interior of the folded protein molecule. Phenylalanine can be converted into tyrosine. |
| Glycine | G | Gly | Because of the two hydrogen atoms at the α carbon, glycine is not optically active. It is the smallest amino acid, rotates easily, and adds flexibility to the protein chain. It is able to fit into the tightest spaces, e.g., the triple helix of collagen. As too much flexibility is usually not desired, as a structural component, it is less common than alanine. |
| Histidine | H | His | His is essential for humans. In even slightly acidic conditions, protonation of the nitrogen occurs, changing the properties of histidine and the polypeptide as a whole. It is used by many proteins as a regulatory mechanism, changing the conformation and behavior of the polypeptide in acidic regions such as the late endosome or lysosome, enforcing conformation change in enzymes. However, only a few histidines are needed for this, so it is comparatively scarce. |
| Isoleucine | I | Ile | Ile is essential for humans. Isoleucine, leucine, and valine have large aliphatic hydrophobic side chains. Their molecules are rigid, and their mutual hydrophobic interactions are important for the correct folding of proteins, as these chains tend to be located inside of the protein molecule. |
| Leucine or isoleucine | J | Xle | A placeholder when either amino acid may occupy a position |
| Lysine | K | Lys | Lys is essential for humans, and behaves similarly to arginine. It contains a long, flexible side chain with a positively charged end. The flexibility of the chain makes lysine and arginine suitable for binding to molecules with many negative charges on their surfaces. E.g., DNA-binding proteins have their active regions rich with arginine and lysine. The strong charge makes these two amino acids prone to be located on the outer hydrophilic surfaces of the proteins; when they are found inside, they are usually paired with a corresponding negatively charged amino acid, e.g., aspartate or glutamate. |
| Leucine | L | Leu | Leu is essential for humans, and behaves similarly to isoleucine and valine. |
| Methionine | M | Met | Met is essential for humans. Always the first amino acid to be incorporated into a protein, it is sometimes removed after translation. Like cysteine, it contains sulfur, but with a methyl group instead of hydrogen. This methyl group can be activated, and is used in many reactions where a new carbon atom is being added to another molecule. |
| Asparagine | N | Asn | Similar to aspartic acid, Asn contains an amide group where Asp has a carboxyl. |
| Pyrrolysine | O | Pyl | Similar to lysine, but it has a pyrroline ring attached. |
| Proline | P | Pro | Pro contains an unusual ring to the N-end amine group, which forces the CO-NH amide sequence into a fixed conformation. It can disrupt protein folding structures like α helix or β sheet, forcing the desired kink in the protein chain. Common in collagen, it often undergoes a post-translational modification to hydroxyproline. |
| Glutamine | Q | Gln | Similar to glutamic acid, Gln contains an amide group where Glu has a carboxyl. Used in proteins and as a storage for ammonia, it is the most abundant amino acid in the body. |
| Arginine | R | Arg | Functionally similar to lysine. |
| Serine | S | Ser | Serine and threonine have a short group ended with a hydroxyl group. Its hydrogen is easy to remove, so serine and threonine often act as hydrogen donors in enzymes. Both are very hydrophilic, so the outer regions of soluble proteins tend to be rich with them. |
| Threonine | T | Thr | Essential for humans, Thr behaves similarly to serine. |
| Selenocysteine | U | Sec | The selenium analog of cysteine, in which selenium replaces the sulfur atom. |
| Valine | V | Val | Essential for humans, Val behaves similarly to isoleucine and leucine. |
| Tryptophan | W | Trp | Essential for humans, Trp behaves similarly to phenylalanine and tyrosine. It is a precursor of serotonin and is naturally fluorescent. |
| Unknown | X | Xaa | Placeholder when the amino acid is unknown or unimportant. |
| Tyrosine | Y | Tyr | Tyr behaves similarly to phenylalanine (precursor to tyrosine) and tryptophan, and is a precursor of melanin, epinephrine, and thyroid hormones. Naturally fluorescent, its fluorescence is usually quenched by energy transfer to tryptophans. |
| Glutamic acid or glutamine | Z | Glx | A placeholder when either amino acid may occupy a position |

Catabolism
Amino acids can be classified according to the properties of their main products:[13]
- Glucogenic, with the products having the ability to form glucose by gluconeogenesis
- Ketogenic, with the products not having the ability to form glucose: These products may still be used for ketogenesis or lipid synthesis.
- Amino acids catabolized into both glucogenic and ketogenic products
See also
References
- ^ Ambrogelly A, Palioura S, Söll D (January 2007). "Natural expansion of the genetic code". Nature Chemical Biology. 3 (1): 29–35. doi:10.1038/nchembio847. PMID 17173027.
- ^ Lobanov AV, Turanov AA, Hatfield DL, Gladyshev VN (August 2010). "Dual functions of codons in the genetic code". Critical Reviews in Biochemistry and Molecular Biology. 45 (4): 257–65. doi:10.3109/10409231003786094. PMC 3311535. PMID 20446809.
- ^ Young VR (August 1994). "Adult amino acid requirements: the case for a major revision in current recommendations" (PDF). The Journal of Nutrition. 124 (8 Suppl): 1517S–1523S. doi:10.1093/jn/124.suppl_8.1517S. PMID 8064412.
- ^ Erives A (August 2011). "A model of proto-anti-codon RNA enzymes requiring L-amino acid homochirality". Journal of Molecular Evolution. 73 (1–2): 10–22. Bibcode:2011JMolE..73...10E. doi:10.1007/s00239-011-9453-4. PMC 3223571. PMID 21779963.
- ^ Frenkel-Pinter, Moran; Haynes, Jay W.; C, Martin; Petrov, Anton S.; Burcar, Bradley T.; Krishnamurthy, Ramanarayanan; Hud, Nicholas V.; Leman, Luke J.; Williams, Loren Dean (2019-08-13). "Selective incorporation of proteinaceous over nonproteinaceous cationic amino acids in model prebiotic oligomerization reactions". Proceedings of the National Academy of Sciences. 116 (33): 16338–16346. Bibcode:2019PNAS..11616338F. doi:10.1073/pnas.1904849116. ISSN 0027-8424. PMC 6697887. PMID 31358633.
- ^ Thurlkill RL, Grimsley GR, Scholtz JM, Pace CN (May 2006). "pK values of the ionizable groups of proteins". Protein Science. 15 (5): 1214–8. doi:10.1110/ps.051840806. PMC 2242523. PMID 16597822.
- ^ Pace CN, Grimsley GR, Scholtz JM (May 2009). "Protein ionizable groups: pK values and their contribution to protein stability and solubility". The Journal of Biological Chemistry. 284 (20): 13285–9. doi:10.1074/jbc.R800080200. PMC 2679426. PMID 19164280.
- ^ Byun BJ, Kang YK (May 2011). "Conformational preferences and pK(a) value of selenocysteine residue". Biopolymers. 95 (5): 345–53. doi:10.1002/bip.21581. PMID 21213257. S2CID 11002236.
- ^ Rother M, Krzycki JA (August 2010). "Selenocysteine, pyrrolysine, and the unique energy metabolism of methanogenic archaea". Archaea. 2010: 1–14. doi:10.1155/2010/453642. PMC 2933860. PMID 20847933.
- ^ Kozlowski LP (January 2017). "Proteome-pI: proteome isoelectric point database". Nucleic Acids Research. 45 (D1): D1112–D1116. doi:10.1093/nar/gkw978. PMC 5210655. PMID 27789699.
- ^ "Atomic Weights and Isotopic Compositions for All Elements". NIST. Retrieved 2016-12-12.
- ^ Phillips R, Kondev J, Theriot J, Garcia HG, Orme N (2013). Physical biology of the cell (Second ed.). Garland Science. p. 178. ISBN 978-0-8153-4450-6.
- ^ Ferrier DR (2005). "Chapter 20: Amino Acid Degradation and Synthesis". In Champe PC, Harvey RA, Ferrier DR (eds.). Lippincott's Illustrated Reviews: Biochemistry (Lippincott's Illustrated Reviews). Hagerstwon, MD: Lippincott Williams & Wilkins. ISBN 978-0-7817-2265-0.
General references
- Nelson, David L.; Cox, Michael M. (2000). Lehninger Principles of Biochemistry (3rd ed.). Worth Publishers. ISBN 978-1-57259-153-0.
- Kyte J, Doolittle RF (May 1982). "A simple method for displaying the hydropathic character of a protein". Journal of Molecular Biology. 157 (1): 105–32. CiteSeerX 10.1.1.458.454. doi:10.1016/0022-2836(82)90515-0. PMID 7108955.
- Meierhenrich, Uwe J. (2008). Amino acids and the asymmetry of life (1st ed.). Springer. ISBN 978-3-540-76885-2.
- Biochemistry, Harpers (2015). Harpers Illustrated Biochemistry (30st ed.). Lange. ISBN 978-0-07-182534-4.