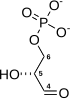
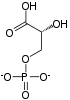
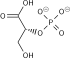
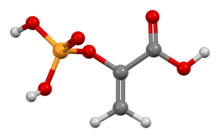
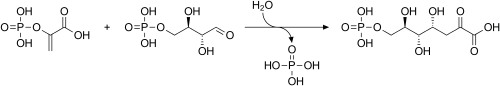Phosphoenolpyruvic acid
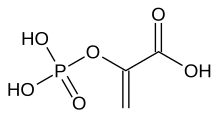
| |
| |
| Names | |
|---|---|
| Preferred IUPAC name 2-(Phosphonooxy)prop-2-enoic acid | |
| Other names Phosphoenolpyruvic acid, PEP | |
| Identifiers | |
3D model (JSmol) |
|
| ChEBI | |
| ChemSpider | |
| DrugBank | |
| ECHA InfoCard | 100.004.830 |
| KEGG | |
PubChem CID |
|
| UNII | |
CompTox Dashboard (EPA) |
|
| |
| |
| Properties | |
| C3H5O6P | |
| Molar mass | 168.042 |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
Phosphoenolpyruvate (2-phosphoenolpyruvate, PEP) is the carboxylic acid derived from the enol of pyruvate and phosphate. It exists as an anion. PEP is an important intermediate in biochemistry. It has the highest-energy phosphate bond found (−61.9 kJ/mol) in organisms, and is involved in glycolysis and gluconeogenesis. In plants, it is also involved in the biosynthesis of various aromatic compounds, and in carbon fixation; in bacteria, it is also used as the source of energy for the phosphotransferase system.[1][2]
In glycolysis
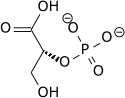
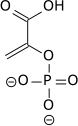
PEP is formed by the action of the enzyme enolase on 2-phosphoglyceric acid. Metabolism of PEP to pyruvic acid by pyruvate kinase (PK) generates adenosine triphosphate (ATP) via substrate-level phosphorylation. ATP is one of the major currencies of chemical energy within cells.
| 2-phospho-D-glycerate | Enolase | phosphoenolpyruvate | Pyruvate kinase | pyruvate | ||
|
|

| ||||
| H2O | ADP | ATP | ||||
|
| |||||
| H2O | ||||||
Compound C00631 at KEGG Pathway Database. Enzyme 4.2.1.11 at KEGG Pathway Database. Compound C00074 at KEGG Pathway Database. Enzyme 2.7.1.40 at KEGG Pathway Database. Compound C00022 at KEGG Pathway Database.
In gluconeogenesis
PEP is formed from the decarboxylation of oxaloacetate and hydrolysis of one guanosine triphosphate molecule. This reaction is catalyzed by the enzyme phosphoenolpyruvate carboxykinase (PEPCK). This reaction is a rate-limiting step in gluconeogenesis:[3]
- GTP + oxaloacetate → GDP + phosphoenolpyruvate + CO2
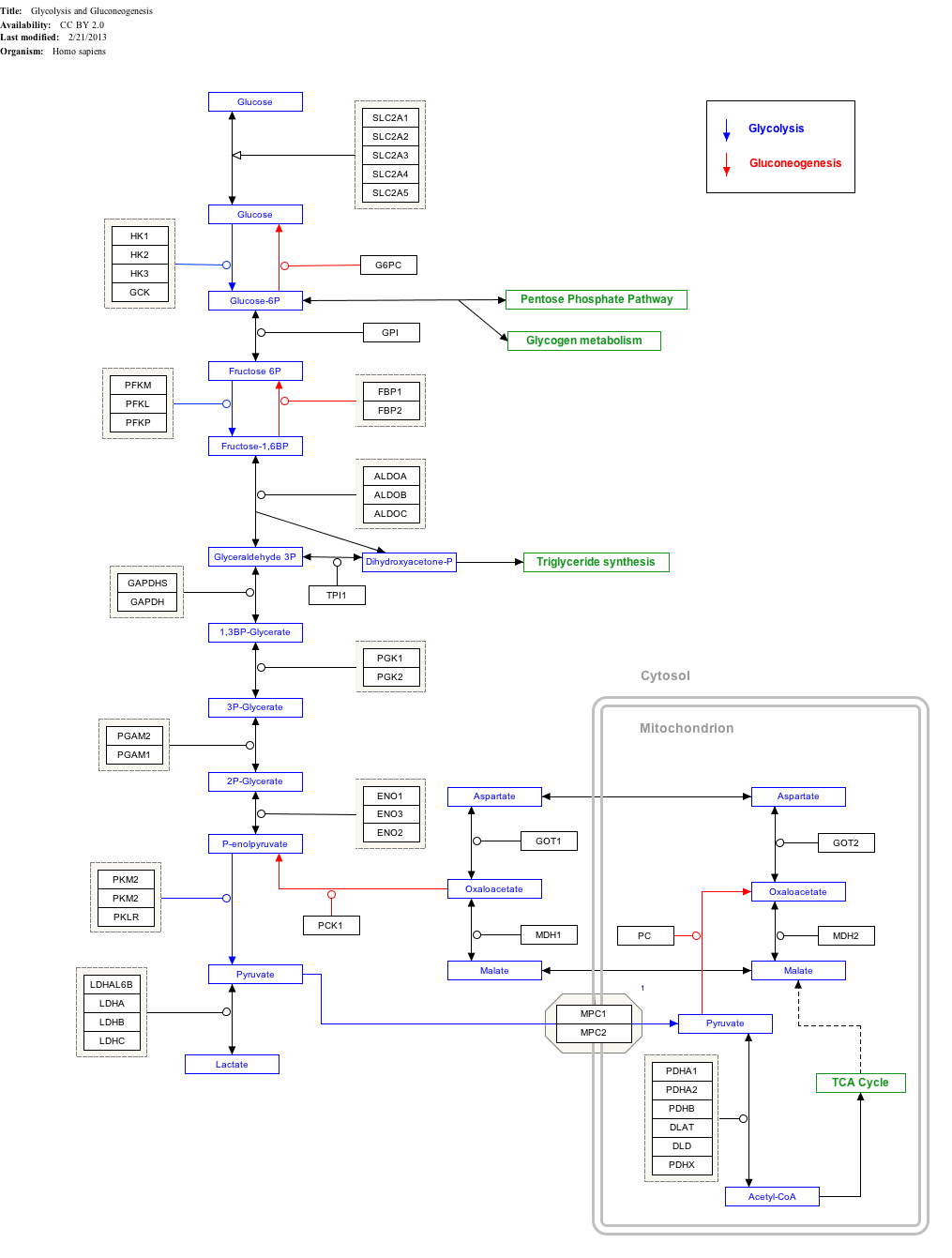
Interactive pathway map
Click on genes, proteins and metabolites below to link to respective articles.[§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "GlycolysisGluconeogenesis_WP534".
In plants
PEP may be used for the synthesis of chorismate through the shikimate pathway.[4] Chorismate may then be metabolized into the aromatic amino acids (phenylalanine, tryptophan and tyrosine) and other aromatic compounds. The first step is when Phosphoenolpyruvate and erythrose-4-phosphate react to form 3-deoxy-D-arabinoheptulosonate-7-phosphate (DAHP), in a reaction catalyzed by the enzyme DAHP synthase.
In addition, in C4 plants, PEP serves as an important substrate in carbon fixation. The chemical equation, as catalyzed by phosphoenolpyruvate carboxylase (PEP carboxylase), is:
- PEP + HCO−3 → oxaloacetate
References
- ^ Berg, Jeremy M.; Tymoczko, Stryer (2002). Biochemistry (5th ed.). New York: W.H. Freeman and Company. ISBN 0-7167-3051-0.
- ^ Nelson, D. L.; Cox, M. M. "Lehninger, Principles of Biochemistry" 3rd Ed. Worth Publishing: New York, 2000. ISBN 1-57259-153-6.
- ^ "InterPro: IPR008209 Phosphoenolpyruvate carboxykinase, GTP-utilising". Retrieved 2007-08-17.
- ^ "BioCarta - Charting Pathways of Life". Retrieved 2007-08-17.