Inward-rectifier potassium channel
| Inward rectifier potassium channel | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 crystal structure of an inward rectifier potassium channel | |||||||||
| Identifiers | |||||||||
| Symbol | IRK | ||||||||
| Pfam | PF01007 | ||||||||
| Pfam clan | CL0030 | ||||||||
| InterPro | IPR013521 | ||||||||
| SCOP2 | 1n9p / SCOPe / SUPFAM | ||||||||
| TCDB | 1.A.2 | ||||||||
| OPM superfamily | 8 | ||||||||
| OPM protein | 3SPG | ||||||||
| |||||||||
Inward-rectifier potassium channels (Kir, IRK) are a specific lipid-gated subset of potassium channels. To date, seven subfamilies have been identified in various mammalian cell types,[1] plants,[2] and bacteria.[3] They are activated by phosphatidylinositol 4,5-bisphosphate (PIP2). The malfunction of the channels has been implicated in several diseases.[4][5] IRK channels possess a pore domain, homologous to that of voltage-gated ion channels, and flanking transmembrane segments (TMSs). They may exist in the membrane as homo- or heterooligomers and each monomer possesses between 2 and 4 TMSs. In terms of function, these proteins transport potassium (K+), with a greater tendency for K+ uptake than K+ export.[3] The process of inward-rectification was discovered by Denis Noble in cardiac muscle cells in 1960s[6] and by Richard Adrian and Alan Hodgkin in 1970 in skeletal muscle cells.[7]
Overview of inward rectification
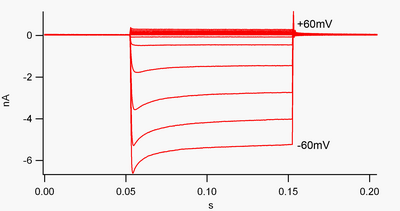
A channel that is "inwardly-rectifying" is one that passes current (positive charge) more easily in the inward direction (into the cell) than in the outward direction (out of the cell). It is thought that this current may play an important role in regulating neuronal activity, by helping to stabilize the resting membrane potential of the cell.
By convention, inward current (positive charge moving into the cell) is displayed in voltage clamp as a downward deflection, while an outward current (positive charge moving out of the cell) is shown as an upward deflection. At membrane potentials negative to potassium's reversal potential, inwardly rectifying K+ channels support the flow of positively charged K+ ions into the cell, pushing the membrane potential back to the resting potential. This can be seen in figure 1: when the membrane potential is clamped negative to the channel's resting potential (e.g. -60 mV), inward current flows (i.e. positive charge flows into the cell). However, when the membrane potential is set positive to the channel's resting potential (e.g. +60 mV), these channels pass very little current. Simply put, this channel passes much more current in the inward direction than the outward one, at its operating voltage range. These channels are not perfect rectifiers, as they can pass some outward current in the voltage range up to about 30 mV above resting potential.
These channels differ from the potassium channels that are typically responsible for repolarizing a cell following an action potential, such as the delayed rectifier and A-type potassium channels. Those more "typical" potassium channels preferentially carry outward (rather than inward) potassium currents at depolarized membrane potentials, and may be thought of as "outwardly rectifying." When first discovered, inward rectification was named "anomalous rectification" to distinguish it from outward potassium currents.[8]
Inward rectifiers also differ from tandem pore domain potassium channels, which are largely responsible for "leak" K+ currents.[9] Some inward rectifiers, termed "weak inward rectifiers", carry measurable outward K+ currents at voltages positive to the K+ reversal potential (corresponding to, but larger than, the small currents above the 0 nA line in figure 1). They, along with the "leak" channels, establish the resting membrane potential of the cell. Other inwardly rectifying channels, termed "strong inward rectifiers," carry very little outward current at all, and are mainly active at voltages negative to the K+ reversal potential, where they carry inward current (the much larger currents below the 0 nA line in figure 1).[10]
Mechanism of inward rectification
The phenomenon of inward rectification of Kir channels is the result of high-affinity block by endogenous polyamines, namely spermine, as well as magnesium ions, that plug the channel pore at positive potentials, resulting in a decrease in outward currents. This voltage-dependent block by polyamines results in efficient conduction of current only in the inward direction. While the principal idea of polyamine block is understood, the specific mechanisms are still controversial.[11]
Activation by PIP2
All Kir channels require phosphatidylinositol 4,5-bisphosphate (PIP2) for activation.[12] PIP2 binds to and directly activates Kir 2.2 with agonist-like properties.[13] In this regard Kir channels are PIP2 ligand-gated ion channels.
Role
Kir channels are found in multiple cell types, including macrophages, cardiac and kidney cells, leukocytes, neurons, and endothelial cells. By mediating a small depolarizing K+ current at negative membrane potentials, they help establish resting membrane potential, and in the case of the Kir3 group, they help mediate inhibitory neurotransmitter responses, but their roles in cellular physiology vary across cell types:
| Location | Function |
|---|---|
| cardiac myocytes | Kir channels close upon depolarization, slowing membrane repolarization and helping maintain a more prolonged cardiac action potential. This type of inward-rectifier channel is distinct from delayed rectifier K+ channels, which help repolarize nerve and muscle cells after action potentials; and potassium leak channels, which provide much of the basis for the resting membrane potential. |
| endothelial cells | Kir channels are involved in regulation of nitric oxide synthase. |
| kidneys | Kir export surplus potassium into collecting tubules for removal in the urine, or alternatively may be involved in the reuptake of potassium back into the body. |
| neurons and in heart cells | G-protein activated IRKs (Kir3) are important regulators, modulated by neurotransmitters. A mutation in the GIRK2 channel leads to the weaver mouse mutation. "Weaver" mutant mice are ataxic and display a neuroinflammation-mediated degeneration of their dopaminergic neurons.[14] Relative to non-ataxic controls, Weaver mutants have deficits in motor coordination and changes in regional brain metabolism.[15] Weaver mice have been examined in labs interested in neural development and disease for over 30 years. |
| pancreatic beta cells | KATP channels (composed of Kir6.2 and SUR1 subunits) control insulin release. |
Regulation
Voltage-dependence may be regulated by external K+, by internal Mg2+, by internal ATP and/or by G-proteins. The P domains of IRK channels exhibit limited sequence similarity to those of the VIC family. Inward rectifiers play a role in setting cellular membrane potentials, and closing of these channels upon depolarization permits the occurrence of long duration action potentials with a plateau phase. Inward rectifiers lack the intrinsic voltage sensing helices found in many VIC family channels. In a few cases, those of Kir1.1a, Kir6.1 and Kir6.2, for example, direct interaction with a member of the ABC superfamily has been proposed to confer unique functional and regulatory properties to the heteromeric complex, including sensitivity to ATP. These ATP-sensitive channels are found in many body tissues. They render channel activity responsive to the cytoplasmic ATP/ADP ratio (increased ATP/ADP closes the channel). The human SUR1 and SUR2 sulfonylurea receptors (spQ09428 and Q15527, respectively) are the ABC proteins that regulate both the Kir6.1 and Kir6.2 channels in response to ATP, and CFTR (TC #3.A.1.208.4) may regulate Kir1.1a.[16]
Structure
The crystal structure[17] and function[18] of bacterial members of the IRK-C family have been determined. KirBac1.1, from Burkholderia pseudomallei, is 333 amino acyl residues (aas) long with two N-terminal TMSs flanking a P-loop (residues 1-150), and the C-terminal half of the protein is hydrophilic. It transports monovalent cations with the selectivity: K ≈ Rb ≈ Cs ≫ Li ≈ Na ≈ NMGM (protonated N-methyl-D-glucamine). Activity is inhibited by Ba2+, Ca2+, and low pH.[18]
Classification
There are seven subfamilies of Kir channels, denoted as Kir1 – Kir7.[1] Each subfamily has multiple members (i.e. Kir2.1, Kir2.2, Kir2.3, etc.) that have nearly identical amino acid sequences across known mammalian species.
Kir channels are formed from as homotetrameric membrane proteins. Each of the four identical protein subunits is composed of two membrane-spanning alpha helices (M1 and M2). Heterotetramers can form between members of the same subfamily (i.e. Kir2.1 and Kir2.3) when the channels are overexpressed.
Diversity
| Gene | Protein | Aliases | Associated subunits |
|---|---|---|---|
| KCNJ1 | Kir1.1 | ROMK1 | NHERF2 |
| KCNJ2 | Kir2.1 | IRK1 | Kir2.2, Kir4.1, PSD-95, SAP97, AKAP79 |
| KCNJ12 | Kir2.2 | IRK2 | Kir2.1 and Kir2.3 to form heteromeric channel, auxiliary subunit: SAP97, Veli-1, Veli-3, PSD-95 |
| KCNJ4 | Kir2.3 | IRK3 | Kir2.1 and Kir2.3 to form heteromeric channel, PSD-95, Chapsyn-110/PSD-93 |
| KCNJ14 | Kir2.4 | IRK4 | Kir2.1 to form heteromeric channel |
| KCNJ3 | Kir3.1 | GIRK1, KGA | Kir3.2, Kir3.4, Kir3.5, Kir3.1 is not functional by itself |
| KCNJ6 | Kir3.2 | GIRK2 | Kir3.1, Kir3.3, Kir3.4 to form heteromeric channel |
| KCNJ9 | Kir3.3 | GIRK3 | Kir3.1, Kir3.2 to form heteromeric channel |
| KCNJ5 | Kir3.4 | GIRK4 | Kir3.1, Kir3.2, Kir3.3 |
| KCNJ10 | Kir4.1 | Kir1.2 | Kir4.2, Kir5.1, and Kir2.1 to form heteromeric channels |
| KCNJ15 | Kir4.2 | Kir1.3 | |
| KCNJ16 | Kir5.1 | BIR 9 | |
| KCNJ8 | Kir6.1 | KATP | SUR2B |
| KCNJ11 | Kir6.2 | KATP | SUR1, SUR2A, and SUR2B |
| KCNJ13 | Kir7.1 | Kir1.4 |
Diseases related to Kir channels
- Persistent hyperinsulinemic hypoglycemia of infancy is related to autosomal recessive mutations in Kir6.2. Certain mutations of this gene diminish the channel's ability to regulate insulin secretion, leading to hypoglycemia.
- Bartter's syndrome can be caused by mutations in Kir channels. This condition is characterized by the inability of kidneys to recycle potassium, causing low levels of potassium in the body.
- Andersen's syndrome is a rare condition caused by multiple mutations of Kir2.1. Depending on the mutation, it can be dominant or recessive. It is characterized by periodic paralysis, cardiac arrhythmias and dysmorphic features. (See also KCNJ2)
- Barium poisoning is likely due to its ability to block Kir channels.
- Atherosclerosis (heart disease) may be related to Kir channels. The loss of Kir currents in endothelial cells is one of the first known indicators of atherogenesis (the beginning of heart disease).
- Thyrotoxic hypokalaemic periodic paralysis has been linked to altered Kir2.6 function.[19]
- EAST/SeSAME syndrome is caused by mutations in KCNJ10.[20]
See also
References
- ^ a b Kubo Y, Adelman JP, Clapham DE, Jan LY, Karschin A, Kurachi Y, et al. (December 2005). "International Union of Pharmacology. LIV. Nomenclature and Molecular Relationships of Inwardly Rectifying Potassium Channels". Pharmacological Reviews. 57 (4): 509–26. doi:10.1124/pr.57.4.11. PMID 16382105. S2CID 11588492.
- ^ Hedrich R, Moran O, Conti F, Busch H, Becker D, Gambale F, et al. (1995). "Inward rectifier potassium channels in plants differ from their animal counterparts in response to voltage and channel modulators". European Biophysics Journal. 24 (2): 107–15. doi:10.1007/BF00211406. PMID 8582318. S2CID 12718513.
- ^ a b "1.A.2 Inward Rectifier K Channel (IRK-C) Family". TCDB. Retrieved 2016-04-09.
- ^ Hansen SB (May 2015). "Lipid agonism: The PIP2 paradigm of ligand-gated ion channels". Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids. 1851 (5): 620–8. doi:10.1016/j.bbalip.2015.01.011. PMC 4540326. PMID 25633344.
- ^ Abraham MR, Jahangir A, Alekseev AE, Terzic A (November 1999). "Channelopathies of inwardly rectifying potassium channels". FASEB Journal. 13 (14): 1901–10. doi:10.1096/fasebj.13.14.1901. PMID 10544173. S2CID 22205168.
- ^ Noble, Denis (December 1965). "Electrical properties of cardiac muscle attributable to inward going (anomalous) rectification". Journal of Cellular and Comparative Physiology. 66 (S2): 127–135. doi:10.1002/jcp.1030660520. ISSN 0095-9898.
- ^ Adrian RH, Chandler WK, Hodgkin AL (July 1970). "Slow changes in potassium permeability in skeletal muscle". The Journal of Physiology. 208 (3): 645–68. doi:10.1113/jphysiol.1970.sp009140. PMC 1348790. PMID 5499788.
- ^ Bertil Hille (2001). Ion Channels of Excitable Membranes 3rd ed. (Sinauer: Sunderland, MA), p. 151. ISBN 0-87893-321-2.
- ^ Hille, p. 155.
- ^ Hille, p. 153.
- ^ Lopatin AN, Makhina EN, Nichols CG (November 1995). "The mechanism of inward rectification of potassium channels: "long-pore plugging" by cytoplasmic polyamines". The Journal of General Physiology. 106 (5): 923–55. doi:10.1085/jgp.106.5.923. PMC 2229292. PMID 8648298.
- ^ Tucker SJ, Baukrowitz T (May 2008). "How highly charged anionic lipids bind and regulate ion channels". The Journal of General Physiology. 131 (5): 431–8. doi:10.1085/jgp.200709936. PMC 2346576. PMID 18411329.
- ^ Hansen SB, Tao X, MacKinnon R (August 2011). "Structural basis of PIP2 activation of the classical inward rectifier K+ channel Kir2.2". Nature. 477 (7365): 495–8. Bibcode:2011Natur.477..495H. doi:10.1038/nature10370. PMC 3324908. PMID 21874019.
- ^ Peng J, Xie L, Stevenson FF, Melov S, Di Monte DA, Andersen JK (November 2006). "Nigrostriatal dopaminergic neurodegeneration in the weaver mouse is mediated via neuroinflammation and alleviated by minocycline administration". The Journal of Neuroscience. 26 (45): 11644–51. doi:10.1523/JNEUROSCI.3447-06.2006. PMC 6674792. PMID 17093086.
- ^ Strazielle C, Deiss V, Naudon L, Raisman-Vozari R, Lalonde R (October 2006). "Regional brain variations of cytochrome oxidase activity and motor coordination in Girk2(Wv) (Weaver) mutant mice". Neuroscience. 142 (2): 437–49. doi:10.1016/j.neuroscience.2006.06.011. PMID 16844307. S2CID 33064439.
- ^ WO application 0190360, Wei MH, Chaturvedi K, Guegler K, Webster M, Ketchum KA, Di Francesco V, Beasley E, "Isolated human transporter proteins, nucleic acid molecules encoding human transporter proteins, and uses thereof", published 29 November 2001, assigned to Apperla Corporation
- ^ Kuo A, Gulbis JM, Antcliff JF, Rahman T, Lowe ED, Zimmer J, et al. (June 2003). "Crystal structure of the potassium channel KirBac1.1 in the closed state". Science. 300 (5627): 1922–6. Bibcode:2003Sci...300.1922K. doi:10.1126/science.1085028. PMID 12738871. S2CID 2703162.
- ^ a b Enkvetchakul D, Bhattacharyya J, Jeliazkova I, Groesbeck DK, Cukras CA, Nichols CG (November 2004). "Functional characterization of a prokaryotic Kir channel". The Journal of Biological Chemistry. 279 (45): 47076–80. doi:10.1074/jbc.C400417200. PMC 8629170. PMID 15448150.
- ^ Ryan DP, da Silva MR, Soong TW, Fontaine B, Donaldson MR, Kung AW, et al. (January 2010). "Mutations in potassium channel Kir2.6 cause susceptibility to thyrotoxic hypokalemic periodic paralysis". Cell. 140 (1): 88–98. doi:10.1016/j.cell.2009.12.024. PMC 2885139. PMID 20074522.
- ^ Bockenhauer D, Feather S, Stanescu HC, Bandulik S, Zdebik AA, Reichold M, et al. (May 2009). "Epilepsy, ataxia, sensorineural deafness, tubulopathy, and KCNJ10 mutations". The New England Journal of Medicine. 360 (19): 1960–70. doi:10.1056/NEJMoa0810276. PMC 3398803. PMID 19420365.
Further reading
External links
- Inward+Rectifier+Potassium+Channels at the U.S. National Library of Medicine Medical Subject Headings (MeSH).
- "Inwardly Recifying Potassium Channels". IUPHAR Database of Receptors and Ion Channels. International Union of Basic and Clinical Pharmacology.
- UMich Orientation of Proteins in Membranes families/family-85 - Spatial positions of inward rectifier potassium channels in membranes.
